false 0001631574 0001631574 2023-12-15 2023-12-15
UNITED STATES
SECURITIES AND EXCHANGE COMMISSION
Washington, D.C. 20549
Form 8-K
CURRENT REPORT
Pursuant to Section 13 or 15(d)
of the Securities Exchange Act of 1934
Date of Report (Date of earliest event reported): December 15, 2023
WAVE LIFE SCIENCES LTD.
(Exact name of registrant as specified in its charter)
|
|
|
|
|
| Singapore |
|
001-37627 |
|
98-1356880 |
(State or other jurisdiction
of incorporation) |
|
(Commission File Number) |
|
(IRS Employer
Identification No.) |
|
|
|
| 7 Straits View #12-00, Marina One East Tower Singapore |
|
|
|
018936 |
| (Address of principal executive offices) |
|
|
|
(Zip Code) |
Registrant’s telephone number, including area code: +65 6236 3388
Check the appropriate box below if the Form 8-K filing is intended to simultaneously satisfy the filing obligation of the registrant under any of the following provisions (see General Instruction A.2. below):
| ☐ |
Written communications pursuant to Rule 425 under the Securities Act (17 CFR 230.425) |
| ☐ |
Soliciting material pursuant to Rule 14a-12 under the Exchange Act (17 CFR 240.14a-12) |
| ☐ |
Pre-commencement communications pursuant to Rule 14d-2(b) under the Exchange Act (17 CFR 240.14d-2(b)) |
| ☐ |
Pre-commencement communications pursuant to Rule 13e-4(c) under the Exchange Act (17 CFR 240.13e-4(c)) |
Indicate by check mark whether the registrant is an emerging growth company as defined in Rule 405 of the Securities Act of 1933 (§230.405 of this chapter) or Rule 12b-2 of the Securities Exchange Act of 1934 (§240.12b-2 of this chapter).
Emerging growth company ☐
If an emerging growth company, indicate by check mark if the registrant has elected not to use the extended transition period for complying with any new or revised financial accounting standards provided pursuant to Section 13(a) of the Exchange Act. ☐
Securities registered pursuant to Section 12(b) of the Act:
|
|
|
|
|
| Title of each class |
|
Trading symbol |
|
Name of each exchange on
which registered |
| $0 Par Value Ordinary Shares |
|
WVE |
|
The Nasdaq Global Market |
| Item 7.01 |
Regulation FD Disclosure. |
From time to time, Wave Life Sciences Ltd. (the “Company”) presents and/or distributes slides and presentations to the investment community to provide updates and summaries of its business. On December 15, 2023, the Company updated its corporate presentation, which is available on the “For Investors & Media” section of the Company’s website at http://ir.wavelifesciences.com/. This presentation is also furnished as Exhibit 99.1 to this Current Report on Form 8-K.
The information in this Item 7.01 and exhibit 99.1 attached hereto is being furnished and shall not be deemed “filed” for purposes of Section 18 of the Securities Exchange Act of 1934, as amended (the “Exchange Act”), or otherwise subject to the liabilities of that Section, nor shall it be deemed incorporated by reference into any registration statement or other filing under the Securities Act of 1933, as amended, or the Exchange Act, except as shall be expressly set forth by specific reference in such filing.
| Item 9.01 |
Financial Statements and Exhibits. |
The following exhibit relating to Item 7.01 is furnished and not filed:
SIGNATURES
Pursuant to the requirements of the Securities Exchange Act of 1934, the registrant has duly caused this report to be signed on its behalf by the undersigned hereunto duly authorized.
|
|
|
| WAVE LIFE SCIENCES LTD. |
|
|
| By: |
|
/s/ Paul B. Bolno, M.D. |
|
|
Paul B. Bolno, M.D. |
|
|
President and Chief Executive Officer |
Date: December 15, 2023

Wave Life Sciences Corporate
Presentation December 15, 2023 Exhibit 99.1

Forward-looking statements This
document contains forward-looking statements. All statements other than statements of historical facts contained in this document, including statements regarding possible or assumed future results of operations, preclinical and clinical studies,
business strategies, research and development plans, collaborations and partnerships, regulatory activities and timing thereof, competitive position, potential growth opportunities, use of proceeds and the effects of competition are forward-looking
statements. These statements involve known and unknown risks, uncertainties and other important factors that may cause the actual results, performance or achievements of Wave Life Sciences Ltd. (the “Company”) to be materially different
from any future results, performance or achievements expressed or implied by the forward-looking statements. In some cases, you can identify forward-looking statements by terms such as “may,” “will,” “should,”
“expect,” “plan,” “aim,” “anticipate,” “could,” “intend,” “target,” “project,” “contemplate,” “believe,” “estimate,”
“predict,” “potential” or “continue” or the negative of these terms or other similar expressions. The forward-looking statements in this presentation are only predictions. The Company has based these
forward-looking statements largely on its current expectations and projections about future events and financial trends that it believes may affect the Company’s business, financial condition and results of operations. These forward-looking
statements speak only as of the date of this presentation and are subject to a number of risks, uncertainties and assumptions, including those listed under Risk Factors in the Company’s Form 10-K and other filings with the SEC, some of which
cannot be predicted or quantified and some of which are beyond the Company’s control. The events and circumstances reflected in the Company’s forward-looking statements may not be achieved or occur, and actual results could differ
materially from those projected in the forward-looking statements. Moreover, the Company operates in a dynamic industry and economy. New risk factors and uncertainties may emerge from time to time, and it is not possible for management to predict
all risk factors and uncertainties that the Company may face. Except as required by applicable law, the Company does not plan to publicly update or revise any forward-looking statements contained herein, whether as a result of any new information,
future events, changed circumstances or otherwise.

Building a leading RNA medicines
company $20 million milestone earned under GSK collaboration and $100 million offering in December 2023 extended cash runway into 4Q 2025* DMD (splicing), HD (silencing), and AATD (RNA editing) clinical programs advancing INHBE, obesity
(siRNA), muscle sparing, fat loss, improved metabolic profile Multi-modal drug discovery and development platform Leader in RNA editing with potential best-in-class oligonucleotide chemistry Strategic collaborations to expand and
advance pipeline In-house GMP manufacturing; Strong and broad IP portfolio DMD, HD, and AATD clinical programs advancing Upcoming Milestones: Proof-of-mechanism data from RestorAATion clinical program of WVE-006 for AATD in 2024 Select INHBE
clinical candidate for metabolic disorders, including obesity, in 4Q 2024 and submit CTA in 2025 Data from FORWARD-53 clinical trial of WVE-N531 for DMD in 2024 Data from SELECT-HD clinical trial of WVE-003 for HD in 2Q 2024 *Cash runway
does not include potential future milestones or opt-in payments under GSK and Takeda collaborations

Combining potential best-in-class
chemistry with novel biology and genetic insights: Opportunities for new high impact medicines Accessing new endogenous enzymes for novel modalities (RNA editing) Opening up new targets, including prevalent diseases

Wave's versatile RNA medicines
platform unlocks genetic insights for rare and common diseases opening up new target opportunities Claussnitzer, et al. Nature (2020) 577, 179; King et al. PLoS Genet (2019) 15, e1008489 Accessing UK Biobank and building proprietary
machine learning models to generate unique genetic insights

Program Discovery Preclinical Clinical
Rights Patient population (US & Europe) RNA EDITING WVE-006 SERPINA1 (AATD) GSK exclusive global license 200K Multiple undisclosed Correction 100% global >20K (multiple) Multiple undisclosed Upregulation 100% global >3M (multiple)
SILENCING: siRNA INHBE* (Metabolic disorders, including obesity) 100% global 47M SPLICING WVE-N531 Exon 53 (DMD) 100% global 2.3K Other exons (DMD) 100% global Up to 18K SILENCING: ANTISENSE WVE-003 mHTT (HD) Takeda 50:50 Option 25K Manifest (SNP3)
60K Pre-Manifest (SNP3) Robust RNA medicines pipeline including first-in-class RNA editing programs FORWARD-53 Trial (Phase 2) SELECT-HD Trial (Phase 1b/2a) RestorAATion Clinical Program *Through GSK collaboration, Wave can advance up to three
collaboration programs (the first of which is INHBE) and GSK can advance up to eight collaboration programs. AATD: Alpha-1 antitrypsin deficiency; DMD: Duchenne muscular dystrophy; HD: Huntington’s disease Editing for correction Editing for
upregulation

Collaboration leverages Wave’s
unique stereopure, PN-chemistry containing PRISMTM platform, including editing, splicing, silencing (RNAi and antisense) Strategic collaboration with GSK to develop transformative RNA medicines for genetically defined diseases 1$120 million in cash
and $50 million equity investment received in January 2023, 2Initiation, development, launch, and commercialization milestones for WVE-006 and programs progressed during initial 4-year research term (8 GSK collaboration programs), 3GSK eligible
to receive tiered royalty payments and commercial milestones from Wave First-in-class RNA editing program GSK granted exclusive global license to WVE-006 for AATD GSK to advance up to eight collaboration programs Up to $225 million in
development and launch milestones Up to $1.2 billion in aggregate in initiation, development and launch milestones Up to $300 million in sales-related milestones Up to $1.6 billion in aggregate in sales-related milestones Double-digit tiered
royalties as a percentage of net sales up to high-teens Tiered royalties as a percentage of net sales up to low-teens Development and commercialization responsibilities transfer to GSK after completion of first-in-patient study Development and
commercialization responsibilities transfer to GSK at development candidate Wave to advance up to three wholly owned collaboration programs (or more pending agreement with GSK) 3 Wave to leverage GSK’s genetic insights Multiple value drivers
to Wave Milestone / royalties Genetic targets Milestone / royalties $170 million upfront to Wave (cash and equity1) Additional research support funding Potential for up to $3.3 billion in milestones2 Expands Wave’s pipeline INHBE is
Wave’s first wholly-owned program emerging from GSK collaboration

WVE-006 (RNA editing) AATD

3) Retain M-AAT physiological
regulation 2) Reduce Z-AAT protein aggregation in liver WVE-006: Designed to correct mutant SERPINA1 transcript to address both liver and lung manifestations of AATD M-AAT reaches lungs to protect from proteases M-AAT secretion into bloodstream AAT:
Alpha-1 antitrypsin Strnad et al., 2020 N Engl J Med 382:1443-55; Blanco et al., 2017 Int J Chron Obstruct Pulmon Dis 12:561-69; Remih et al., 2021 Curr Opin Pharmacol 59:149-56. WVE-006 ADAR editing approach to address key goals of AATD treatment:
RNA correction replaces mutant Z-AAT protein with wild-type M-AAT protein Z-AAT 1) Restore circulating, functional wild-type M-AAT I(G) A SERPINA1 Z allele mRNA encodes Z-AAT protein with E342K mutation Edited SERPINA1 mRNA enables wild-type M-AAT
protein production WVE-006 (GalNAc-conjugated AIMer) WVE-006 designed to correct Z allele mRNA to enable M-AAT protein to be produced 200,000 Pi*ZZ patients in US and Europe

WVE-006 in AATD: First-in-class RNA
editing clinical candidate Potentially comprehensive approach to address both lung and liver manifestations of AATD Increased AAT protein in NSG-PiZ mice Demonstrated functionality of M-AAT protein Confirmed restored wild-type M-AAT protein WVE-006
treatment results in serum AAT protein levels of up to 30 uM in NSG-PiZ mice Overall percentages of serum AAT protein isoforms in NSG-PiZ mice (Week 13) Serum neutrophil elastase inhibition activity in NSG-PiZ mice AATD: Alpha-1 antitrypsin
deficiency; M-AAT protein: wild-type AAT protein; WVE-006 administered subcutaneously (10 mg/kg bi-weekly) in 7-week old NSG-PiZ mice (n=5 per group); Loading dose: 3 x 10 mg/kg at Day 0. Left: Liver biopsies collected at wk 13 (1 wk after last
dose) and SERPINA1 editing quantified by Sanger sequencing; Right: Total serum AAT protein quantified by ELISA; Stats: Two-Way ANOVA with adjustment for multiple comparisons (Tukey) Potent and durable editing yields functional AAT
protein

WVE-006 decreases lobular
inflammation and PAS-D globule size, prevents increase in hepatocyte turnover Left (Lobular inflammation) and Middle (Mitoses): Scatter plot showing inflammation grade or mitoses score. Each circle represents an individual mouse, (Mean ±
SEM); Right (PAS-D Globule Size): 40 largest globules in each of 5 mice were measured. Each circle represents a single PAS-D globule, (Mean ± SEM). Baseline: week 0 (7 weeks old); Treated week 13 (20 weeks old); Stats: Kruskal-Wallis followed
by Dunn’s test Mitoses (NSG PiZ mice, week 13) Fibrosis à Cirrhosis à Hepatocellular Carcinoma Correction of gain-of-function liver phenotypes Lobular inflammation (NSG PiZ mice, week 13) Week 0 Week 13 Week 0 Week 13 Week 0 Week 13
PAS-D-positive globule size (NSG PiZ mice, week 13)

RNA editing only detected at PiZ
mutation site in SERPINA1 transcript (mouse liver) RNA editing across transcriptome (mouse liver) AIMer-directed editing is highly specific in mice SERPINA1 (PiZ mutation site) % Editing Dose 3x10 mg/kg (days 0, 2, 4) SC with AATD AIMer (SA1 –
4). Liver biopsies day 7. RNA-seq to quantify on-target SERPINA1 editing, to quantify off-target editing reads mapped to entire mouse genome; plotted circles represent sites with LOD>3 (N=4), SERPINA1 edit site is indicated No bystander
editing observed on SERPINA1 transcript Coverage Coverage Editing site (PiZ mutation) PBS AATD AIMer C 0% T 100% C 48.2% T 51.8%

Dosing underway in RestorAATion
clinical program; proof of mechanism data in patients with AATD expected in 2024 Dose escalation Study key objectives Safety and tolerability Pharmacokinetics Serum M-AAT levels Dosing Underway Multiple assessments of serum AAT throughout cohort HV:
healthy volunteer; SAD: single-ascending dose; MAD: multi-ascending dose RestorAATion-2: AATD Patients SAD à MAD cohorts Dose E Dose D Dose C Dose B Dose A High dose Medium dose Low dose Informs dose & dose frequency RestorAATion-1:
Healthy Volunteers Up to 7 doses

AIMers RNA editing
capability

The AIMer-targetable
‘Edit-Verse’ is substantial The Edit-verse is the editable gene-disease universe, including upregulation >13,000 genes with a high-probability1 of being amenable to transcriptional regulation with A-to-G editing Model development
ongoing to expand access to more protein-coding genes and expand the Edit-verse AIMers are expected to be able to target ~50% of the transcriptome 1(score >95th p-tile) Gene-Disease Network

Multiple RNA editing opportunities
to build high-value pipeline beyond WVE-006 The Edit-verse is substantial and still expanding Advancing work for a diverse set of undisclosed targets addressing areas of high unmet need, including both rare and prevalent diseases Hepatic
(GalNAc-AIMers) Extra-Hepatic (AIMers) Target A Target B Target X Target E Target F Target G Approach Upregulation Upregulation Upregulation Correction Upregulation Correction Tissue Liver Liver Liver Liver Kidney Lung Therapeutic Area Metabolic
Metabolic Renal Rare Renal Rare Estimated Patients (US and Europe) ~90M ~3M ~170K ~17K ~85K ~5K Potential to advance any combination of targets into preclinical development

INHBE program (siRNA silencing)
Metabolic disorders, including obesity

siRNA silencing is one of multiple
Wave modalities being advanced in strategic research collaboration with GSK Potential for best-in-class siRNA enabled by Wave’s PRISM platform **** Left, Middle, and right: Mice expressing human HSD17B13 transgene treated with siRNA (3 mg/kg)
or PBS, liver mRNA, guide strand concentration, Ago2 loading quantified. Stats: Two-way ANOVA with post-hoc test * P<0.05, ****P<0.0001. Liu et al., 2023 Nuc Acids Res doi: 10.1093/nar/gkad268; Wk 2 Wk 14 Wk 7 Reference Wave siRNA * *
Ago2 loading (liver, transgenic mice) Wave siRNA Reference PBS Unprecedented Ago2 loading increases potency and durability of silencing following administration of single subcutaneous dose 1 Wk 2 Wk 14 Wk 7 1 PBS HSD-1933 HSD-1930 NTC PBS HSD-1933
HSD-1930 PBS HSD-1933 HSD-1930

INHBE: Evolution in treatment
for obesity; muscle sparing, sustained fat loss, improved metabolic profile 1. Liang, et al. 2023 Postgraduate Medical Journal 99(1175):985; 2. Lakka, et al. 2002 JAMA 288(21):2709; 3. Sargeant, et al. 2019 Endocrinol Metab (Seoul)
34(3):247-262; 4. Liu, et al. 2022 Front. Endocrinol. 13:1043789; 5. Prime Therapeutics Claims Analysis, July 2023; 6. Müller, et al. 2019 Molecular Metabolism 30: 72-130. Metabolic syndrome* is associated with type 2 diabetes, cardiovascular
disease, hypertension, stroke, cancer, and increased mortality1,2 Estimate ~47M people in US and Europe with metabolic disorders, including obesity Therapeutic options beyond GLP-1s are needed GLP-1 receptor agonists lead to weight loss at the
expense of muscle mass3 GLP-1 receptor agonists suppress general reward system6 GLP-1 receptor agonists associated with poor tolerability profile4 with 68% drop-off after 1 year5 Preferred approach would improve metabolism and increase fat loss
while maintaining muscle mass Restoration of metabolic health via INHBE silencing expected to simultaneously address obesity and other drivers of metabolic syndrome *Patients diagnosed with metabolic syndrome based on having 3 of the following:
abdominal obesity, high bp, high blood glucose, high TG, or low HDL

Driven by clinical genetics,
GalNac-siRNA program addresses high unmet need in metabolic disorders, including obesity Nat Commun 2022. https://doi.org/10.1038/s41467-022-32398-7; 2. Nat Commun 2022. https://doi.org/10.1038/s41467-022-31757-8; 3. PLOS ONE 2018.
https://doi.org/10.1371/journal.pone.0194798; 4. Adam, RC. et.al. Proc Natl Acad Sci USA. 2023, 120(32): e2309967120. INHBE program is Wave’s first wholly owned program emerging from GSK collaboration Leverages novel genetic insights accessed
through GSK collaboration INHBE loss-of-function heterozygous carriers exhibit healthy metabolic profile1,2,3: Reduced waist-to-hip ratio Reduced odds ratio of type 2 diabetes by 28%, and coronary artery disease Reduced serum triglycerides Elevated
HDL-c Reduced HbA1c Lowered ApoB INHBE expressed primarily in liver and gene product (subunit of activin E) acts on its receptor in adipose tissue4 GalNAc-siRNA for targeted delivery to hepatocytes ≥50% reduction of INHBE with siRNA expected
to restore a healthy metabolic profile

INHBE knockdown of 90% demonstrated
in human hepatocytes with GalNAc-siRNA Primary hepatocytes were treated with a cross-reactive siRNA via free uptake. INHBE mRNA was quantified by RT-qPCR. This cross-reactive sequence demonstrates ~90% maximal knock-down in human hepatocytes and
~65% in mouse hepatocytes Additional human selective sequences are in development Human hepatocytes Mouse hepatocytes

~62% silencing **** Therapeutic
threshold1 INHBE knockdown demonstrated in mice at 5 weeks INHBE silencing achieved in vivo with GalNAc-siRNA exceeds therapeutic threshold and led to lower body weight HFD: high-fat diet. Stats: two-sided Welch’s T Test **** P <
0.0001 1. Adam, RC. et.al. Proc Natl Acad Sci USA. 2023, 120(32): e2309967120. INHBE knockdown led to 16% lower body weight as compared to control Data plotted by body weight difference as a percentage of PBS treated young DIO mice; Coskun, T. et.
al. Mol. Metab. 2018, 18, 3. Stats: Repeated Measures ANOVA; Inhbe siRNA vs. Control significantly different at P < 0.05 level weeks 2 through 5 mRNA expression (relative to PBS liver) Body weight relative to PBS (%) INHBE silencing Lower
relative body weight Control (HFD, PBS) Inhbe siRNA Weeks HFD, PBS Inhbe siRNA Similar effect seen in semaglutide preclinical studies

~56% reduction ~34% reduction ~38%
reduction **** *** *** Chow, PBS HFD, PBS HFD, Inhbe siRNA Chow, PBS HFD, PBS HFD, Inhbe siRNA Chow, PBS HFD, PBS HFD, Inhbe siRNA INHBE silencing leads to significant decrease in visceral fat, consistent with UK Biobank human data
on loss-of-function carriers Adam, RC. et.al. Proc Natl Acad Sci USA. 2023, 120(32): e2309967120. HFD: high-fat diet. Stats: white-adjusted Two-way ANOVA with Bonferroni-adjusted post hoc comparisons per tissue type allowing heteroscedasticity
(only HFD, Inhbe siRNA vs. HFD, PBS shown) *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 mesenteric epididymal inguinal Changes in white adipose tissue after 5 weeks INHBE knockdown in young DIO mice resulted in less fat mass
across multiple types of white adipose tissue, without loss of brown fat Subsequent 8-week study demonstrates further reduction in excess visceral fat Changes in white adipose tissue after 8 weeks mesenteric epididymal inguinal ~56% reduction ~40%
reduction ~45% reduction **** ** * Chow, PBS HFD, PBS HFD, Inhbe siRNA Chow, PBS HFD, PBS HFD, Inhbe siRNA Chow, PBS HFD, PBS HFD, Inhbe siRNA

Foster, DJ. et.al. Mol Ther. 2018,
26(3), 708. B6 mice administered PBS or 0.5 mg/kg of siRNA (subcutaneous). Benchmark: Stats: Mixed Two-way ANOVA followed by post hoc test comparing siRNA vs. Next gen siRNA per day derived from linear mixed effects model * P < 0.0001
Wave’s next generation GalNAc-siRNA demonstrates best-in-class potential Next generation siRNA results in more potent and durable knockdown of serum Ttr protein Next generation siRNA INHBE program Applying next-generation siRNA chemistry to
INHBE program Potent and highly specific leads identified Potential for infrequent administration INHBE candidate for metabolic disorders, including obesity, expected in 4Q 2024; CTA expected in 2025 * * *

Wave’s platform chemistry
enables siRNA extra-hepatic delivery Chemical impact Introduction of neutral backbone Unique structural feature of PN, specifically guanidine Increased lipophilicity Stereochemistry Extra-hepatic delivery Titrating siRNA lipophilicity tunable
PNs (PN variants) Maintaining high Ago2 loading and intracellular trafficking Titrating plasma protein binding Altered delivery, enhanced potency and durability in various tissues PN PN can tune extra-hepatic delivery of siRNA using rational design,
including placement, number of modifications and PN variants

Tunable PN variants enhance potency
and alter extra-hepatic delivery of non-GalNAc siRNAs Stats: Three-way ANOVA followed by Bonferroni-adjusted post hoc test comparing condition to PBS (data not shown) * P < 0.05, *** P < 0.001, **** P < 0.0001; B6 mice
administered PBS or 5 mg/kg of Sod1 siRNA (no GalNAc conjugate) subcutaneous injection (n=7). Taqman qPCR assays used for RNA PD, relative fold changes of Sod1 to Hprt mRNA normalized to % of PBS group. Non-GalNAc siRNA with PN variants
improve silencing in liver and adipose tissue 14 and 28 days post single dose Reaching adipose tissue in addition to liver with siRNA is important for certain metabolic disorders PN variants also enhanced siRNA silencing in muscle tissue,
including heart and diaphragm **** **** **** *** **** **** * Day

Single dose of next generation
siRNA delivers broad, potent and durable CNS target engagement PBS (dotted line) or 100 μg of App siRNA administered ICV (n=7). PCR assays for RNA PD, relative fold changes of App to Hprt mRNA normalized to % of PBS; Stats: Three-way ANOVA
followed by Bonferroni-adjusted post hoc test comparing condition to PBS (data not shown), Next gen siRNA significantly lower than PBS at both time points for all tissues at P < 0.0001 level; Immunohistochemical analysis of FFPE Mouse Brain
tissue labeling App protein (Color Brown) with CS#19389 followed by a ready to use Polymer-HRP 2nd Detection antibody. Nuclei were counterstained with Hematoxylin (Color Blue). Single 100 ug ICV injection Sustained APP knockdown of at least 75%
throughout the 16-week study in vivo in mice APP silencing PBS Next gen siRNA Robust target engagement translates to substantial App protein reduction across brain regions 8-weeks post single dose

Wave siRNA demonstrates more potent
and durable silencing as compared to published state-of-the-art PBS (dotted line) or 100 μg of App siRNA administered ICV (n=7). PCR assays for RNA PD, relative fold changes of App to Hprt mRNA normalized to % of PBS; Stats: Three-way ANOVA
followed by Bonferroni-adjusted post hoc test comparing condition to PBS (data not shown), Next gen siRNA significantly lower than PBS at both time points for all tissues at P < 0.0001 level. Source: Brown, K.M., Nair, J.K., Janas, M.M. et
al. Expanding RNAi therapeutics to extrahepatic tissues with lipophilic conjugates. Nat Biotechnol 40, 1500–1508 (2022). Knockdown < 90 days post-dose Single dose 100 μg by ICV Single dose 120 μg by ICV Alnylam (APP
– Cortex) Wave (APP – Cortex) Nat Biotechnol 40, 1500–1508 (2022) Knockdown > 112 days post-dose %mRNA remaining (SEM) (App/Hprt) % message remaining (relative to aCSF)

WVE-N531 (splicing) Duchenne
muscular dystrophy
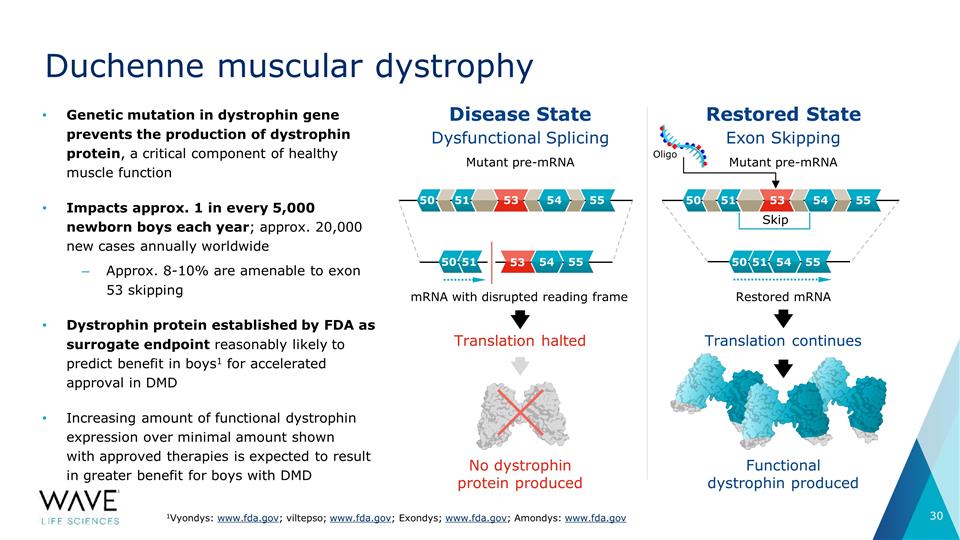
Duchenne muscular dystrophy Genetic
mutation in dystrophin gene prevents the production of dystrophin protein, a critical component of healthy muscle function Impacts approx. 1 in every 5,000 newborn boys each year; approx. 20,000 new cases annually worldwide Approx. 8-10% are
amenable to exon 53 skipping Dystrophin protein established by FDA as surrogate endpoint reasonably likely to predict benefit in boys1 for accelerated approval in DMD Increasing amount of functional dystrophin expression over minimal amount shown
with approved therapies is expected to result in greater benefit for boys with DMD 1Vyondys: www.fda.gov; viltepso; www.fda.gov; Exondys; www.fda.gov; Amondys: www.fda.gov Dysfunctional Splicing Exon Skipping No dystrophin protein produced
Functional dystrophin produced Translation halted Translation continues Mutant pre-mRNA Disease State Restored State mRNA with disrupted reading frame Restored mRNA Mutant pre-mRNA Skip Oligo 53 53 50 51 54 55 50 51 54 55 53 50 51 54 55 50 51 54 55

Extended survival in dKO
preclinical model supports potential of exon-skipping therapeutics for DMD Kandasamy et al., 2022; doi: 10.1093/nar/gkac018 PN chemistry improved function and survival in dKO mice 100% survival at time of study termination Restored muscle and
respiratory function to wild-type levels Note: Untreated, age-matched mdx mice had 100% survival at study termination [not shown] Time (weeks) PS/PO/PN 150 mg/kg weekly PS/PO/PN 75 mg/kg bi-weekly PS/PO 150 mg/kg weekly PBS Survival probability (%)
Tidal volume Age (days) TVb (ml) Wild-type dKO: PBS dKO: PS/PO/PN Wild-type dKO: PBS dKO (PS/PO/PN oligonucleotide)

Preclinical data supported
advancing WVE-N531 to clinical development: enhanced delivery and high levels of dystrophin 26th Annual ASGCT meeting, May 16-20, 2023 WVE-N531 reached high concentrations in heart and diaphragm in NHP WVE-N531: Dystrophin restoration of up to 71%
in vitro Western Blot normalized to primary healthy human myoblast lysate Conc (uM) % Dystrophin Dystrophin Vinculin

Clinical data from WVE-N531 Part A:
High exon-skipping & muscle concentrations after three bi-weekly doses

Dosing underway in FORWARD-53, a
potentially registrational Phase 2 clinical trial of WVE-N531 in DMD (Exon 53) Design of FORWARD-53: Phase 2, open-label, 10 mg/kg every other week, 10 patients enrolled Endpoints: Dystrophin (powered for >5% of normal), safety/tolerability,
pharmacokinetics, digital and functional assessments (incl. NSAA and others) Muscle biopsies to assess dystrophin expression Fully enrolled and dosing underway Screening Safety Follow-up Every other week IV dosing Functional assessment Biopsy after
24 weeks of treatment Functional assessment Biopsy after 48 weeks of treatment Functional assessment IV: intravenous; NSAA: North star ambulatory assessment Potentially registrational dystrophin expression data are expected in 2024

Potential for Wave to address up to
40% of DMD population Exon 45 Exon 44 Exon 52 WVE-N531 Exon 53 Exon 51 Not Amenable to Skipping 11-13% 8-10% 44% Left: Aartsma-Rus, et al. 2009 Hum Mutat 30, 293. DMD Population Exon 52 Exon 51 Exon 44 Protein Restoration Exon skipping and
dystrophin restoration demonstrated in vitro Exon Skipping

WVE-003 (Allele-selective antisense
silencing) Huntington’s Disease

Healthy individual
Huntington’s disease mHTT toxic effects lead to neurodegeneration; loss of wtHTT functions may also contribute to HD Stresses wtHTT Stresses wtHTT mHTT + ~50% decrease in wtHTT Healthy CNS function Synaptic dysfunction | Cell death |
Neurodegeneration Loss of wtHTT functions Huntington’s disease (HD) Wild-type HTT (wtHTT) is critical for normal neuronal function Expanded CAG triplet repeat in HTT gene results in production of mutant huntingtin protein (mHTT) HD is a
monogenic autosomal dominant genetic disease; fully penetrant and affects entire brain Fatal disease characterized by cognitive decline, psychiatric illness, and chorea 30,000 people with HD in the US and more than 200,000 at risk of developing
HD

WVE-003 (SNP3) demonstrates
selective, potent, and durable reduction of mHTT in preclinical models Selectively reduces mHTT mRNA in HD iPSC neurons in vitro Results from ND50036 iPSC-derived medium spiny neurons. Total HTT knockdown quantified by qPCR and normalized to
HPRT1. Oligonucleotide or PBS [100 μg ICV injections through cannula on days 1, 3, 5] delivered to BACHD transgenic. Mean ± SD (n=8, *P<0.0332, ***P<0.0002, ****P<0.0001 versus PBS unless otherwise noted). HPRT1,
hypoxanthine-guanine phosphoribosyl transferase; iPSC, induced pluripotent stem cell; ICV, intracerebroventricular; PBS, phosphate-buffered saline Similar results in cortex Pan-silencing reference compound WVE-003 PBS Weeks *** ****
**** **** **** **** Pan-silencing reference compound WVE-003 Percentage HTT mRNA Remaining Durable striatal mHTT knockdown for 12 weeks in BACHD mouse model

Preservation of wtHTT WVE-003:
First-in-class allele-selective candidate for HD mHTT protein levels Placebo WVE-003 (30 and 60 mg pooled*) wtHTT protein levels Reductions in mean CSF mHTT and preservation of wtHTT observed in pooled analysis of single-dose cohorts in SELECT-HD
clinical study Single dose of WVE-003 Single dose of WVE-003 Reduction in mHTT protein: 22% from baseline 35% vs. placebo mHTT: mutant huntingtin protein; wtHTT: wild-type huntingtin protein *Pooled considering no apparent dose response between 2
cohorts; Data cut-off: August 29, 2022 Data from 30 mg multi-dose cohort with extended follow-up, along with all single-dose data expected 2Q 2024 Reductions in mHTT

Anticipated upcoming
milestones

Anticipated upcoming milestones
AATD: Alpha-1 antitrypsin deficiency; DMD: Duchenne muscular dystrophy; HD: Huntington’s disease; mHTT: Mutant huntingtin; wtHTT: Wild-type huntingtin WVE-006 (AATD) Most advanced RNA editing candidate & potential best-in-class approach
for AATD 2024: Deliver proof-of-mechanism data from RestorAATion clinical program INHBE Program (Metabolic disorders, including obesity) Driven by clinical genetics, with potential to be next-generation therapeutic for obesity 4Q 2024: Select INHBE
clinical candidate 2025: Submit a clinical trial application (CTA) WVE-N531 (DMD) Potential best-in-class approach with highest exon skipping reported 2024: Deliver potentially registrational dystrophin expression data from FORWARD-53 WVE-003 (HD)
First-in-class mHTT lowering, wtHTT-sparing approach 2Q 2024: Deliver data from 30 mg multi-dose cohort with extended follow up, along with all single-dose data Discovery Pipeline & Collaborations Advance collaboration activities with GSK, with
potential for additional cash inflows in 2024 and beyond Select five new clinical candidates by year-end 2025, including INHBE

Wave is poised for significant and
sustained growth Note: Bubble size illustrative of size of total addressable US market (assuming 100% share of addressable patients) AATD WVE-006 DMD WVE-N531 Exon 53 HD WVE-003 SNP3 INHBE metabolic disorders, including obesity Clinical candidate
expected 4Q 2024; CTA in 2025 Four additional clinical candidates by year-end 2025 Value of US Total Addressable Market (TAM) ~$12B >$65B

For more information:
InvestorRelations@wavelifesci.com Thank you!
v3.23.3
| X |
- DefinitionBoolean flag that is true when the XBRL content amends previously-filed or accepted submission.
| Name: |
dei_AmendmentFlag |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionFor the EDGAR submission types of Form 8-K: the date of the report, the date of the earliest event reported; for the EDGAR submission types of Form N-1A: the filing date; for all other submission types: the end of the reporting or transition period. The format of the date is YYYY-MM-DD.
| Name: |
dei_DocumentPeriodEndDate |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:dateItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionThe type of document being provided (such as 10-K, 10-Q, 485BPOS, etc). The document type is limited to the same value as the supporting SEC submission type, or the word 'Other'.
| Name: |
dei_DocumentType |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:submissionTypeItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionAddress Line 1 such as Attn, Building Name, Street Name
| Name: |
dei_EntityAddressAddressLine1 |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionAddress Line 2 such as Street or Suite number
| Name: |
dei_EntityAddressAddressLine2 |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- Definition
+ References
+ Details
| Name: |
dei_EntityAddressCityOrTown |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionISO 3166-1 alpha-2 country code.
| Name: |
dei_EntityAddressCountry |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:countryCodeItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionCode for the postal or zip code
| Name: |
dei_EntityAddressPostalZipCode |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionA unique 10-digit SEC-issued value to identify entities that have filed disclosures with the SEC. It is commonly abbreviated as CIK. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection b-2
| Name: |
dei_EntityCentralIndexKey |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:centralIndexKeyItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionIndicate if registrant meets the emerging growth company criteria. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection b-2
| Name: |
dei_EntityEmergingGrowthCompany |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionCommission file number. The field allows up to 17 characters. The prefix may contain 1-3 digits, the sequence number may contain 1-8 digits, the optional suffix may contain 1-4 characters, and the fields are separated with a hyphen.
| Name: |
dei_EntityFileNumber |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:fileNumberItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionTwo-character EDGAR code representing the state or country of incorporation.
| Name: |
dei_EntityIncorporationStateCountryCode |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:edgarStateCountryItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionThe exact name of the entity filing the report as specified in its charter, which is required by forms filed with the SEC. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection b-2
| Name: |
dei_EntityRegistrantName |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionThe Tax Identification Number (TIN), also known as an Employer Identification Number (EIN), is a unique 9-digit value assigned by the IRS. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection b-2
| Name: |
dei_EntityTaxIdentificationNumber |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:employerIdItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionLocal phone number for entity.
| Name: |
dei_LocalPhoneNumber |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:normalizedStringItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionBoolean flag that is true when the Form 8-K filing is intended to satisfy the filing obligation of the registrant as pre-commencement communications pursuant to Rule 13e-4(c) under the Exchange Act. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 13e
-Subsection 4c
| Name: |
dei_PreCommencementIssuerTenderOffer |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionBoolean flag that is true when the Form 8-K filing is intended to satisfy the filing obligation of the registrant as pre-commencement communications pursuant to Rule 14d-2(b) under the Exchange Act. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 14d
-Subsection 2b
| Name: |
dei_PreCommencementTenderOffer |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionTitle of a 12(b) registered security. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection b
| Name: |
dei_Security12bTitle |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:securityTitleItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionName of the Exchange on which a security is registered. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Number 240
-Section 12
-Subsection d1-1
| Name: |
dei_SecurityExchangeName |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:edgarExchangeCodeItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionBoolean flag that is true when the Form 8-K filing is intended to satisfy the filing obligation of the registrant as soliciting material pursuant to Rule 14a-12 under the Exchange Act. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Exchange Act
-Section 14a
-Number 240
-Subsection 12
| Name: |
dei_SolicitingMaterial |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionTrading symbol of an instrument as listed on an exchange.
| Name: |
dei_TradingSymbol |
| Namespace Prefix: |
dei_ |
| Data Type: |
dei:tradingSymbolItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
| X |
- DefinitionBoolean flag that is true when the Form 8-K filing is intended to satisfy the filing obligation of the registrant as written communications pursuant to Rule 425 under the Securities Act. Reference 1: http://www.xbrl.org/2003/role/presentationRef
-Publisher SEC
-Name Securities Act
-Number 230
-Section 425
| Name: |
dei_WrittenCommunications |
| Namespace Prefix: |
dei_ |
| Data Type: |
xbrli:booleanItemType |
| Balance Type: |
na |
| Period Type: |
duration |
|
Wave Life Sciences (NASDAQ:WVE)
過去 株価チャート
から 8 2024 まで 9 2024

Wave Life Sciences (NASDAQ:WVE)
過去 株価チャート
から 9 2023 まで 9 2024
